Using a Taxon Split input as an output
We’ve introduced a new way to use Taxon Splits that gives curators more flexibility. Here’s some background before explaining the change. If this background is already familiar to you jump ahead to the second to last section titled Using the Taxon Split input as the output.
Taxon Swaps
On iNaturalist if you want to change the main scientific name associated with a taxon (e.g. from Species X to Species Y) you do the following steps:
- Create a new inactive taxon with the name Species Y
- Create a draft Taxon Swap with Species X as the input and Species Y as the output
- Commit the Taxon Swap
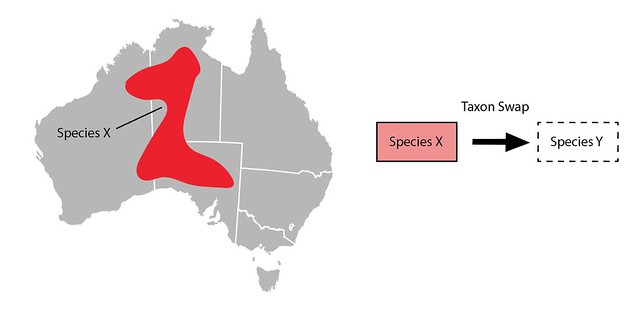
Why do all this instead of just changing the name on the taxon? Just changing the name would mean that if an identifier typed in an identification of Species X it might unexpectedly change to Species Y without any record of what happened. Create a taxon swap leaves a record so identifiers can see when, how, and why the name associated with their identification changed.
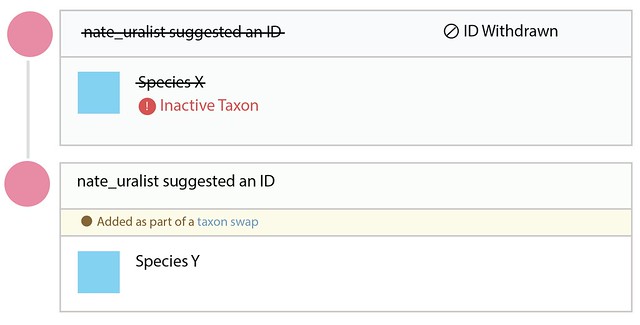
Let’s call this updating identifications. We’re aware that this might not be the easiest, least disruptive way to generate such a log, especially when the name changes are trivial such as going from Cnemaspis hitihami to Cnemaspis hitihamii. But under the current functionality, taxon swaps are always used to change the main scientific name of a taxon.
As shown above, taxon swaps have a single output which means all identifications of the input taxon can be unambiguously updated with identifications of the output taxon. Likewise, other content associated with the input taxon, such as taxon names (where iNaturalist stores common names and synonyms), taxon ranges, listed taxa etc., is moved or copied over to the output taxon*. Committing a taxon swap also makes the input taxon inactive and activates inactive output taxa.
Using Taxon Swaps to Lump Taxa
Another common curation action is to lump one taxon into another (e.g. Species X into Species Z). A taxon swap can also be used to do this by making Species X the input and the already active Species Z as the output. As described above, committing the taxon swap will inactivate Species X, update all identifications, and copy/move other content over to Species Z.
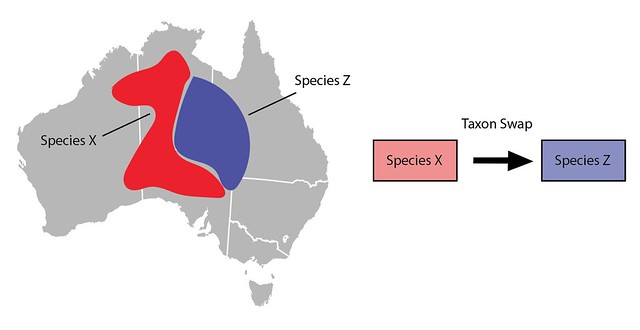
One thing to keep in mind when using Taxon Swaps to lump taxa is that lumping often broadens what we mean by the output taxon. In this example Species Z’s taxon range to the east no longer matches its newly broadened meaning and would need to be manually modified to include the western part of the range. Similarly, imagine Species X’s common name was "Western Critter" and Species Z’s common name was "Eastern Critter". Post-swap, Species’ Z’s common name might need to be updated to something like "Critter".

Several uses for Taxon Merges
If you want to lump two or more taxa (Species X and Species Q) into an existing taxon (Species Z) you can use a Taxon Merge. Taxon Merges work like taxon swaps but have multiple input taxa. Since there’s just a single output taxon, identifications and content are also migrated to the output taxon*.
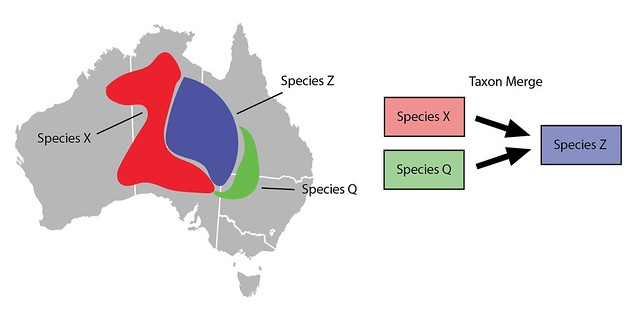
Taxon Merges can also be used to lump taxa slightly differently. Imagine that rather than manually broadening all the content associated with Species Z you prefer to just make a new taxon for Species Z. Let's call it Species Z’. It would start out inactive, would have the same main scientific name as Species Z, but would have a different broader meaning and thus different broader content such as range maps.
Committing the Taxon Merge would inactivate the input Species Z and activate the broader output Species Z', identifications of Species Z would be updated with identifications of Species Z’ and all other content would be copied to Species Z’.
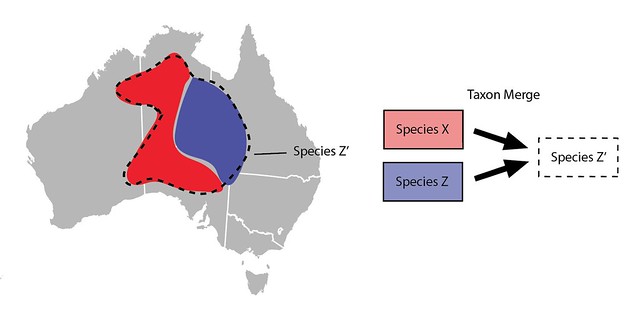
This approach requires more steps than just manually broadening Species Z (Species Z’ must be created etc.) and perhaps unnecessarily updates all the identifications of Species Z with Species Z` (unnecessary since both taxa have the same the main scientific name). So this way of lumping taxa is generally not preferred, but it is an option available to curators.
Taxon Splits
To carve one taxon off from another (e.g. Species W off from Species X) we use Taxon Splits. Taxon splits have a single input and more than one output. The only way to make taxon splits in the past was to create new inactive taxa for the carved-off taxon (Species W) and for the new narrowed version of the input taxon (Species X’).

Since splits have more than one output taxon, identifications are updated to the common ancestor of the outputs. Other content such as range maps and names are not copied over to the output taxa and must be added manually*.

Atlases provide some flexibility for determining a single output taxon with which to update identifications. Using the example of the atlases below, identifications associated with observations in Western Australia (WA) and South Australia (SA) would be updated with output taxon Species X’ while identifications belonging to observations in Queensland (QLD) would be updated with output taxon Species W. Only identifications belonging to observations within presence places in both atlases (e.g. Northern Territory, NT) or outside of both atlases (e.g. New South Wales) would be updated with the common ancestor.

Using the Taxon Split input as the output
Sometimes only a tiny piece of a taxon is carved off as a new taxon. When the input taxon has a lot of identifications this cran result in many unecessary identification updates (e.g. Species X with Species X’).
We’ve made some changes that allow curators the flexibility, if they choose, to make Taxon Splits where the input taxon is also one of the output taxa. In some ways, this approach is analogous to using a taxon swap to broaden an existing taxon (e.g. Species X) instead of using a taxon merge with a new output taxon (e.g. Species X’) as was described above. In this approach, the input taxon is first manually narrowed by modifying the atlas (as well as other content such as range map, etc.) which will be used to determine the destination of identifications.

Upon committing such a Taxon Split, iNaturalist will not inactivate the input taxon. It will still use the atlases to determine which identifications should be updated with which output taxon, but any identification determined to be updated with the output taxon matching the input (e.g. Species X) will not be touched.
Because doing a split this way won’t update as many identifications (e.g. updating identifications of Species X with identifications of Species X’), it will always be less disruptive. But it’s important to remember to manually narrow the input taxon by updating content such as the taxon range and common names that likely no longer match the narrowed meaning taxon after the split.
There are some advantages from the normal way of splitting taxa such as having the inactivated input taxon (Species X) persist alongside the narrowed taxon (Species X’) coexisting in the database for a more thorough comparison of how the taxon was narrowed. However, as iNaturalist has grown, it is our feeling that the disruptive costs both in terms of processing time and in terms of identification clutter of having so many updated identifications (e.g. Species X to Species X’) now outweighs these advantages, especially when there are a lot of identifications on the input taxon (Species X) that are not destined to be carved off (e.g. as Species W). As long as curators remember to manually narrow content such as taxon ranges, we anticipate that this new way of splitting taxa will generally be the preferred way to split taxa moving forward.
Examples
Here are two examples of this new kind of taxon split I committed. The first involved splitting two Caribbean island endemics (Boa orophias on St. Lucia and, Boa nebulosa on Dominica) off from Boa constrictor. Notice that the same Boa constrictor taxon is listed as both the input and an output.

As expected, this observation from St. Lucia was properly updated to Boa orophias whereas this captive observation outside of all three atlases was updated to Genus Boa. And this observation from within the range of Boa constrictor was untouched.
Similarly, this split involved splitting two species (Varanus tsukamotoi from Guam and other Pacific Islands and Varanus bennetti from several other islands including Palau) off from Varanus indicus. Again note that the same taxon Varanus indicus acts as both input and output.

As expected, this observation from Palau was updated as Varanus bennetti, this observation from Guam was updated as Varanus tsukamotoi, and this observation from Papua was untouched.
Note that in both of these examples, I manually narrowed both the taxon range and atlas of the input taxa before committing the splits.
*The details about which content is copied over and which is moved over to the output taxon when a taxon change is committed is complicated. Here’s an attempt to explain exactly what happens. In both a taxon swap and a taxon merge, taxon photos and taxon names are copied over. The copy of the valid scientific name from the input taxon is invalid (a synonym) on the output taxon. Taxon swaps, but not taxon merges, copy the taxon range, conservation statuses, and atlas from the input (unless a taxon range or atlas or conservation status for the same place already exists on the output taxon). Listed taxa are moved/merged from the input taxon to the output taxon for both taxon swaps and taxon merges. For taxon splits, no content is copied over (since the output taxon is ambiguous). Listed taxa are moved to a single output taxon if atlases can be used to uniquely determine one, much like atlases are used to update identifications.





Comments
This is cool. That last option is a nice lower-overhead way to deal with recircumscriptions where existing observations on iNat aren't practically effected.
Nice, iNaturalist just keeps getting better and better!
Super nice!
@loarie Thanks for this update! Question: when a curator sets up this kind of split, did you add any kind of visual reminder (in the taxon change interface) of what the curator will need to check and update for the narrowed taxon? Your asterisked paragraph above covers it pretty well, but it is a long and divergent list to commit to memory, or to remember to find in this blog post. (And, knowing how long the list is, I wonder if it will cause curators to weigh differently the pros and cons of using the new versus old split method?)
jdmore - there are currently no visual reminders about what to check but thats a good idea. But the info in the asterisked paragraph is exactly the same for both kinds of splits - it just different for splits, swaps, and merges
Thanks, and point well taken. I'm thinking the outcome of the automated Listed Taxon changes will be a bit different between the two kinds of splits. But that may be because I don't fully grasp how atlases and listed taxa interact and update each other, especially when atlas places or listed taxa are manually removed, as would likely have to occur for a narrowed taxon.
jdmore - when you "Remove this place" from an atlas page it will destroy all listed taxa on the default checklist for that place or any place descending from that place.
When you commit a split, the atlases are used to determine the output taxon destination for each listed taxon on the input (else the common ancestor). If the output taxon is determined to be the same taxon as the input taxon in this new kind of split - its left untouched. Otherwise its moved to the destination. If a listed taxon already exists for that taxon at that place, the listed taxon is merged into it.
Does that help make sense of it? I don't really think it makes sense for taxon changes to move listed taxa to a taxon that has an atlas - but thats how it works now.
Yes, thanks, that does help!
@loarie does it also remove listings on child checklists ?
So if you remove something from the Brazil primary checklist, will it also remove it from a child checklist such as 'birds of Brazil'?
And what about community curated places, including those that aren't grafted to a standard place?
atlases are only aware of default checklists - so if an atlas place lights up or doesn't its only because of whats on default checklists not other checklists like 'birds of brazil' - similarly if you remove an atlas presence place it will only touch default checklists and won't touch other checklists. But atlases are aware of/do touch the default checklists of descendant places (standard and community curated) - so if there's a listed taxon of the default checklist for a community curated park descending from a country, the country will light up (even if there's no listed taxon for the country). And removing the atlas presence place for that country will destroy the listed taxon on the descending community park default checklist.
Thanks for this explanation. Somehow I was unaware that iNaturalist has such a thing as species ranges. Where can we see that? How are they created? I was aware of the atlas function and had tried to use it but gave up in frustration when I believe it blocked me creating an atlas county by county, said there were too many inputs. Looks like I shouldt try giving the atlas guide (https://www.inaturalist.org/pages/atlases).another look.
@janetwright - any taxon that has a species range defined can be seen on the map tab of the taxon page. Look for the pink range, sometimes it is easier to turn off the other layers to see it.
Creating them requires either obtaining or creating your own KML file that delineates the range and then importing it into the taxon. This obviously means having access to and familiarity with some kind of GIS software package.
If the split taxon is used as an output, are the ungrafted* community curated place listed taxa also destroyed?
*not descending from a standard place
Thanks, @cmcheatle. I can handle kml files, might think about this.
bouteloua - they aren't destroyed but their taxon will be replaced with the common ancestor. For example, this listed taxon on the (ungrafted community curated) Latinamerica and Caribbean Check List was associated with Boa constrictor but after the split is now associated with Boa
https://www.inaturalist.org/listed_taxa/3598550.
One thing thats a little weird though is that the listed taxa of descendant taxa aren't considered. For example this listed taxon of Boa constrictor ssp. constrictor on the same checklist wasn't touched https://www.inaturalist.org/listed_taxa/10009956
@loarie can you make a video tutorial of splitting a taxa ?
Most of the time, looking were you move buttons and tuff make easier to understand.
I want to separate a species in two subspecies but it's a species with two thousands observations, I am worry about create a big mess. On the other hand, I do not want to rename 500 observations one by one...
Friendly,
Jean-Michel.
@jmmaes I don't have bandwidth to make a video in the short term, but I'd be happy to answer specific questions about the split you're trying to do - maybe flag the taxon involved explaining what you're trying to do and mention me and I can help recommend next steps?
This is really great, thanks @loarie!
Thanks @loarie, I will try to do it with a species I can split, where there are not so many specimens, so if does not work, I can repair manually.
Friendly,
Jean-Michel.
@jmmaes what exactly are you trying to split? The only way a species to multiple subspecies split would be valid is if the species itself was determined to be invalid and replaced with multiple newly defined subspecies.
If you are trying to simply add more precise ID's at the subspecies level, then a taxa split is not the appropriate tool for this.
@cmcheatle tahnks. I have a beetle with many observations and many synonyms. Now having many observations, I can see that USA specimens and West coast of Mexico is one subspecies and East coast of Mexico and Central America is another ssp. Mexico and Central America are around 500 observations. USA and West Mexico around 2000. So the idea of setting a ssp. name for a bunch of observations seems save a lot of time. If taxa split is not a good way, is there another way ?
Friendly,
Jean-Michel.
Can you flag the beetle taxon in question and in a comment on it describe the situation and mention me and cmcheatle?
Epilachna mexicana:
https://www.inaturalist.org/taxa/319695-Epilachna-mexicana
total 574 observations.
central american subspecies around 150 observations.
I can do it one by one but would be more fun and fast if I can cut part of them from the map and change name. Than I can recheck those near of the limit for control.
@jmmaes - apologies for any confusion, but maybe good to take the opportunity to clarify here. It sounds like you want to use a taxon change to 'bulk identify' alot of observations from the species rank to the subspecies rank? Taxon changes should never be used to identify species like this - they should only be used to make sure content gets moved to the right spot in response to alterations to the structure of the tree. I know it might seem tedious, but once you add the spp to the tree the proper way to try to identify those observations currently sitting at species Epilachna mexicana to that subspecies is to add identifications.
@loarie - thanks a lot. I was also thinlking that it was too nice to be true. If I have time to play with those, I will do the changes case by case. I have done it for some butterflies but not with so many specimens. Anyway I will change one or two first to see it there is a problem somewhere, kind of a test.
Friendly,
Jean-Michel.
hi jmmaes - if I'm understanding what you mean by "change one or two" as adding your own identifications of subspecies to observations - then that sounds great. As long as you're not using taxon changes as a way to replace other people's identifications of species with identifications of subspecies which should not be done.
@loarie are atlas places supposed to be removed if the peeled-off taxon also has that place? E.g. https://www.inaturalist.org/atlases/29619
no if the peeled-off taxon and the remaining piece of the taxon both occur in a place then both their atlases should contain that place. iNat will respond by rolling IDs in that place back tot he common ancestor
In that example, the IDs were correctly rolled up to the common ancestor because both outputs had Yukon in their atlases, but then somehow Yukon was removed from latifolia. Do you know why?
I'll look into that - it could be a bug - thanks
@loarie I'm confused about the Boa example. You say you moved two subspecies to a new species rank. "Boa constrictor ssp. nebulosa (Dominica) and Boa constrictor ssp. orophias (St. Lucia) elevated to species status." Were these subspecies not recognized by iNaturalist when you made the change? If these subspecies were recognized, why not just swap the subspecies with the species rank? Is the taxon split just for splitting a single taxon based on range? Curious as I will be moving all varieties of a species already on iNaturalist up to the species rank after a paper is published. I'm wondering what the proper way to do that is.
@keirmorse - in the Boa constrictor example you have to do both: (a) swap the ssp into the species and also (b) split the sp into the output species
swap Boa constrictor ssp. nebulosa -> Boa nebulosa
swap Boa constrictor ssp. orophias -> Boa orophias
split Boa constrictor -> Boa constrictor, Boa orophias, Boa nebulosa
the split is necessary to replace existing ID's of Boa constrictor with ID's of the elevated ssp
Happy to help brainstorm through the specific example if you mention me on the taxon change draft
@loarie So the split part was done because there were observations only ID'd to Boa constrictor that needed to go to Boa orophias or Boa nebulosa? Or this tackles specific people's IDs like if one person ID'd to Boa constrictor ssp. nebulosa and another only ID'd the same observation to Boa constrictor as that would create an ID conflict after just a swap of the ssp.?
the former. But the former prevents the latter
@loarie Got it. Thanks!
Add a Comment