Introducing the iNaturalist Geomodel
Today, we’ve changed the Seen Nearby label on suggestions to Expected Nearby. The label comes from predictions made by the iNaturalist Geomodel that we’re introducing for the first time.
What is the Geomodel?
Most of you are familiar with the iNaturalist Computer Vision Model which takes an image as input and returns the most likely species based on visual similarity as an output. We train that model on a set of about 80 thousand species with enough data and update it monthly (we released version 2.7 today).
The iNaturalist Geomodel takes a location as input and returns the most likely species at that location as output. Like the Computer Vision Model, it is a Deep Learning model trained on the same set of taxa and updated on the same monthly schedule. We developed and published the Geomodel in collaboration with the same Visipedia team that assisted with the iNaturalist Computer Vision Model. The map below shows Geomodel predictions for American Pika. The Geomodel is trained only on iNaturalist observations and an elevation map.

From Gridded Observations to Geomodel Predictions
iNaturalist has been using the Geomodel to weight computer vision suggestions since June of the year. We started using the Geomodel to apply the Expected Nearby label today.
Previously, we used a gridded version of the raw observations to weight Computer Vision suggestions and apply the Nearby label. We counted the relative number of observations for each species onto a 1-degree grid. If there were any observations of the taxon in the surrounding 9 grid cells, we applied the Seen Nearby label to suggestions. We used the relative number of observations in the grid cells to weight the Computer Vision suggestions. Note the grid cell for Mexican Treehopper in southern Brazil likely due to a misidentified observation.

We’re now using the Expected Nearby Map predicted by the Geomodel to apply the Nearby label. You can think of the Expected Nearby Map as an estimate of whether the species is present near the location.

The change in name from Seen Nearby to Expected Nearby is intended to make it clear that the label comes from a model prediction rather than a grid of observations. Note that these predictions aren’t perfect. For example, Mexican Treehopper probably doesn’t occur in the Galapagos or Cuba despite the predictions. For some species the Geomodel performs remarkably well, while for others predictions have very high error. Work to better understand these and experiment with improvements is ongoing. But as we show below, on average the Geomodel improves upon the 1-degree grid approach it replaces and we expect continued improvements with future Geomodel versions.
We use an Unthresholded version of the Expected Nearby Map to weight Computer Vision suggestions. You can think of the Unthresholded Map as the relative probability that a species occurs at a location.

You can explore the Expected Nearby Maps and Unthresholded Maps we use to weight Computer vision suggestions on new Geomodel prediction pages we’ve linked from the taxon pages of all approximately included 80,000 species.

Why the Geomodel and Next Steps
We transitioned from the 1-degree gridded data to the Geomodel for four main reasons:
1. Improvements to Computer Vision suggestions
As detailed in the Evaluating the Geomodel section below, the Geomodel improves the accuracy of Computer Vision suggestions compared to the 1-degree grid approach. Version 2.7, released today, is about a 4% improvement over the 1-degree grid approach for Top 1 suggestion accuracy and we anticipate more accuracy gains with future Geomodel versions as we refine the modeling approach and more observations are uploaded.
2. Future direction: Fast/offline geospatial information
The number of Geomodel parameters is less than 2% the size of the 1-degree grid cell data. This means the Geomodel is small and fast enough to run on the mobile device like the Computer Vision Model does on Seek. This opens up the potential for including geospatial information in features such as the Seek in-camera suggestions and displaying taxon maps on mobile devices offline. We haven’t built these features yet, but the Geomodel will make them possible.
3. Future direction: Surfacing unusual observations
As iNaturalist grows, the community needs better tools to surface unusual observations that may represent misidentifications or important discoveries such as a range extension or the early detection of an invasive species.
The figure below shows 2.1 million dragonfly observations ranked by their geographic unusualness as predicted by the Geomodel. The right side of the histogram shows the most unusual 0.01% of observations. We sent these 223 unusual observations to dragonfly expert @dennispaulson to vet. 197 observations (88%) were misidentified observations (red bars) such as this Rainpool Spreadwing misidentified as a Slender Spreadwing. The remaining 26 represented some legitimately unusual records (white bars) such as this Slaty Skimmer range extension from Colorado.

Some observations in the white bars were unusual to our model but not to @dennispaulson, such as this Highland Meadowhawk from Haiti that the Geomodel thinks is unusual. With more observations and identifications from poorly sampled regions, the accuracy of the Geomodel will improve over time.
Fly expert @zdanko helped with a similar experiment with 500,000 hoverfly observations. Similar to dragonflies, of the 365 most unusual observations, 267 observations (73%) were misidentifications.
We’re excited about the potential to build tools around the Geomodel to help more quickly surface these unusual observations for more attention from experts so that misidentifications can be fixed and important discoveries like species range extensions aren’t missed.
4. Future direction: Context about range size
One of the most important characteristics of a species from a conservation perspective is its geographic range size. All other things being equal, small-range species tend to be at much greater risk of extinction than species that are widely distributed. In order to prioritize scarce conservation resources and attention, land managers need tools to determine which species are small-ranged local endemics (species that occur nowhere else in the world) from other more widely distributed species.
As described in the Evaluating the Geomodel section below, Geomodel predictions of range area are well correlated with the areas of range maps such as the Taxon Ranges that appear on some taxon pages that come from external sources.
The figure below shows Geomodel predictions of range area for 10 small-ranged birds from around the world. We hope to build tools around the Geomodel to make it easier to determine which observations belong to small ranged endemic species in order to help the land management community prioritize these conservation targets.

The Expected Nearby Maps are being rendered on the Geomodel prediction pages at a coarse 1.8 thousand square-kilometer resolution and therefore are not publicly revealing precise information about sensitive species. We continue to improve iNaturalist channels that securely mobilize sensitive species data and precise predictions for conservation purposes.
Evaluating the Geomodel
We have evaluated the Geomodel by measuring:
- Improvements to suggestion accuracy
- Retaining the correct suggestion in the Expected Nearby subset
- Overlap between Expected Nearby maps and Taxon Ranges
1. Improvements to suggestion accuracy
On average, Top 1 suggestion accuracy improved from 75% to 83% (+8%) by weighting the raw Computer Vision scores with the 1-degree grid. Weighting with the Geomodel instead improved Top 1 suggestion accuracy to 87% (+12%). We repeated this analysis within geographic and taxonomic groupings and in all cases the Geomodel outperformed the 1-degree data.

2. Retaining the correct suggestion in the Expected Nearby subset
By default, we only show the subset of Nearby suggestions. This has the advantage of removing suggestions that are unlikely based on location, but there’s also a risk of removing the correct suggestion. We calculated Recall statistics measuring how often the correct suggestion was retained in the Nearby subsets derived from the Geomodel and the 1-degree grid. On average, both approaches yielded the same Recall of 0.94 meaning for every 100 observations the correct result was included in the Nearby subset 94 times.

3. Overlap between Expected Nearby maps and Taxon Ranges
To measure how well the Expected Nearby maps compared to the Taxon Ranges displayed on the iNaturalist taxon pages, we compared them and calculated Precision and Recall statistics. The Taxon Ranges aren’t perfectly accurate either so for evaluation purposes we used the subset of around 5,000 Taxon Ranges that contained at least 90% of the observations for the taxon.

We repeated this analysis comparing the 1-degree grids and Geomodel to the Taxon Ranges. The Geomodel predictions improved the average of Precision and Recall. The F1 statistic (the harmonic mean of Precision and Recall) improved by 9% for the Geomodel compared to the 1-degree grid.

The Geomodel also does a better job of matching Taxon Range area than the 1-degree grids as measured by Mean Logarithmic Squared Error (MLSE).

Thank you
We want to extend special thanks to our research collaborators, including Oisin Mac Aodha (University of Edinburgh), Elijah Cole (Caltech), Grant Van Horn (UMass Amherst), Christian Lange (University of Edinburgh), Pietro Perona (Caltech), and @tbrooks (IUCN), as well as the generous support from a Climate Change AI 2021-2022 Innovation Grant that helped make this work possible.
We’re excited about the gains in suggestion accuracy the Geomodel is making possible today and the potential for future directions that it opens up for us to pursue in the coming months. Thank you to the entire iNaturalist community for generating all of the observations and identifications that make training powerful models like the Geomodel possible!





Comments
This is so friggin' exciting! I especially like the capability to view unthresholded maps. If you need someone to review predictions for a few moth groups (e..g Cisthenini or Acentropinae), just let me know!
All very exciting! Thank you folks!
Really exciting new developments!
A stunning piece of work! Looking forward to seeing some of those maps. Thanks for sharing.
This is so unbelievably cool! I hope you realize that I just bombarded my non-science freind with so much excitement about this. Also, out of curiosity, is there a way to view these calculated maps for non-curated species. For example, I was wondering if I could use it for certain gall species that currently do not have a good idea for range.
Exciting stuff.
I presume it is only based on wild records. Curious bleeding of unthreshholded map into coastal areas for Protea cynaroides, and for Expected nearby map in California and se Australia.
https://www.inaturalist.org/geo_model/132848/explain
A quick survey of some of our Cape Flora species, leads me to ask please: how many modelled species are confined to a single hexagon? (e.g. https://www.inaturalist.org/geo_model/566080/explain)
And the curious case or Protea denticulata which is predicted to occur where it does not, but the prediction fails to predict where it actually is:
https://www.inaturalist.org/taxa/574546-Protea-denticulata
& https://www.inaturalist.org/geo_model/574546/explain
it's really great to have developed this, I can't wait for this model to evolve with new environmental variables to take into account
@tonyrebelo we're currently training the Geomodel on all records with a community ID (even captive) which explains why the Protea is showing up in CA and Australia. This is because the main use case right now is to improve Computer Vision suggestions and we haven't had a chance to explore how excluding them would impact existing use-cases where people try to ID garden plants etc.
But I agree that for some of the biogeography future directions it would make more sense to use only wild records - more generally, work needs to be done on thinking about what constitutes a species distribution in the Anthropocene when a species literally has a non-zero probability of showing up anywhere on Earth!
No doubt, this is great and awesome! A great advancement for iNat.
As related, there is a very strong correlation between referenced range maps and identification of entities within those range maps. It is a bit of the tail wagging the dog and should be heavily considered. For example, since much of the iNat crew is based in California and likely all of them have posted a detection of a tree frog (Pseudacris), I don't think anyone can reliably separate morphologically or by any other field characteristic the 3 Pseudacris in the state (if they are still valid entities), but magically nearly all of the identifications match the range map cartooned by Gary Nafis of Californiaherps.com, an exceptional resource. But note, even he says others (USGS) don't believe P. regilla (current taxon) occurs in California.
Keep up the great work. It's really neat to see this come out. I look forward to applying to conservation efforts.
~B
This is great!! I am wondering if there is a hard barrier to expanding Geomodel training data beyond only iNaturalist observations? Seems like the model could be made even more robust by including a selection of relatively well-curated specimen datasets from GBIF. This would bring advantages of a much longer temporal sample, and in many areas of the globe a more complete spatial sample as well.
Very exciting, and glad to see the improvements in accuracy of CV suggestions! I think the potential to "double check" unlikely IDs is great for both finding likely mis-IDed observations, but also really valuable ones that provide novel information - I'm excited to see how that develops.
In terms of interpretation, above it says "You can think of the Unthresholded Map as the relative probability that a species occurs at a location." However, I would think that these maps more likely show the relative probability that a species will be observed (by an iNat user) at a given location, given that the model is produced from iNat data. In cases where there are sparse observations or data deficient species, probability of observation and probability of occurrence might be quite different.
The Geomodel is trained on iNaturalist observations and an elevation map (as an environmental variable). So this is a type of ecological niche model, then. Correct? Will additional environmental variables be added in the future?
Excellent work. Well done! Thank you for the continual improvement of the tools.
Yes, now it should stop suggesting Old World Argiope spp. in Ohio! 😊
@cthawley I agree that the model makes assumptions about species absences that might not be correct, there may be biases that aren't being fully accounted for, and uncertainty/error in those predictions is much large in places that are not well sampled (both errors of commission and errors of omission) - and explicitly modeling probability of being observed separate from probability of occurrence is probably a good future direction to better deal with these biases.
@pfau_tarleton you can read more about the methodology here and here. If you pull out the deep learning location encoding which essentially allows each species to draw on information from all the other species the model collapses to Logistic Regression niche model (LR in Table 1 in the first paper). But a huge part of the strength of this approach as opposed to a single species niche model is that the species learns from all other 80k species being modeled (much like the Computer Vision Model) so the model gets a good sense for co-occurrence, biogeography and the kind of things species distributions tend to do without having to rely so much on environmental covariates alone as a crutch as traditional niche models do. This is why the predictions are pretty good using just elevation as a covariate and not including other typical covariates like precipitation etc. We tested adding those covariates and didn't get significant improvement but made the model more complicated.
I agree this is just a baby step though, lots of avenues for improvement, and different approaches might be needed to push these into other applications and scales. We're focused on improving computer vision suggestions at the moment even though I'm also excited by some of these future directions.
@loarie Ooooooh, I get the picture now. That's really interesting! Thanks!
Awesome! Thanks!
This is hugely exciting, congrats to everyone involved. Adding the preprint to my reading list.
Also thanks for the additional response to @pfau_tarleton, super interesting.
@loarie I just tried the live version of the model on some of my observations and suddenly realized that the predicted range doesn't work well with migratory species: yes, many of the predicted species will occur at my location in 3-4 months from now but they are definitely not "Expected Nearby" at this particular time of the year (and would be extreme rarities if they did). Any plans to include seasonality in future iterations?
''We calculated Recall statistics measuring how often the correct suggestion was retained in the Nearby subsets derived from the Geomodel and the 1-degree grid. On average, both approaches yielded the same Recall of 0.94 meaning for every 100 observations the correct result was included in the Nearby subset 94 times.''
Can the user of Computer Vision predict or expect if the result is incorrect and belongs to the 6 times where the correct result is not included.,eg because iNaturalist does not have many observations on the location of the observation?
----june2024
When the geomodel is used, the one-degree-grid method is not used.
https://forum.inaturalist.org/t/make-observations-nearby-circle-smaller/52256/6
@optilete - if the suggestions don't include the right species, it could be because of errors in the Geomodel resulting from gaps/biases in training data, but could be other sources of error or errors in the Computer Vision Model. If the suggestion is being included in the Nearby subset thats due to errors in the Geomodel perhaps stemming from gaps in the training data or elsewhere

What would cause the geomodel to predict magellanic horned and austral pygmy owls to occur in Antarctica? Is it just elevation?


Magellanic horned owl:
Austral pygmy owl:
@brennafarrell I suspect the model doesn't yet know what to make of Antarctica since there's so little data from there - but if you look at the Unthresholded map there is a threshold that could be drawn that would cause those Antarctica cells to drop out without loosing any/too much of the Patagonia range - which brings up the issue of choosing the correct thresholds which we still haven't perfected as evidence here. It would have ideally computed a more conservative threshold https://www.inaturalist.org/geo_model/1456383/explain
not to get too much ahead of ourselves, but it would be neat to be able to capture the community's expertise on absences and feed that into the model. For example you're saying 'I know Mellagenic Horned Owl doesn't occur here', that's useful information and it would be neat to try to capture that from the community in order to better teach the model - maybe a bit like how atlases are being used on the site currently
Has any consideration been given to including or substituting the Level 1, 2, or 3 Ecoregions (EPA, etc.) as part of the predictive build for the Geomodel for North America? Since they are expressly derived from other fundamental ecological data sets, they would move the predictive basis forward substantially. I find them infinitely more applicable to organism distributions than things like elevation or raw climatic variables.
See, for instance: https://www.epa.gov/eco-research/ecoregions
Of course, they do not constitute a global data set, but I'm wondering if there are equivalent sets for much of the globe.
@gcwarbler That is a great idea! That would definitely help with range.
@brennafarrell Nice point and good example of how data sparse areas can be challenging. Fortunately, there aren't going to be too many observers in Antarctica that might get confused by a suggestion like this! It definitely would be interesting to try to incorporate absence data, especially for larger places that are otherwise suitable, but species haven't been able to disperse there. Since the model uses other species occurrences /biogeography, areas where some members of a community are capable of dispersing, but others aren't might be challenges for accurate predictions in the model leading to overprediction for dispersal limited species.
This is a fantastic upgrade. Looking forward to seeing it develop and smooth out as it progresses.
I am not sure what any of this quite means but it sounds like a tremendous amount of work so congratulations on launching it. Thank you for the work that goes into this site that you make available to us. I also wanted to say thank you for including Membracis mexicana. It made me smile to see a garden friend! (They are quite adorable, like little roosters.)
@loarie Thanks! I was wondering if in the future it would be possible to remove incorrect sections from the geomodel. I also have another question. For the sunda scops owl:

the places that are blue, like Sulawesi, Cambodia, and Thailand, will it be included in the 'Expected Nearby' CV there?
@cthawley Thanks! I'm not too concerned with Antarctica, more with other incorrect places included in the new geomodel.
@brennafarrell, correct the blue is the predicted Expected Nearby Map which drives the Expected Nearby Label which is incorrect (false positive) as evaluated by the independent Taxon Range (pink or purple where there’s overlap) - annoying it’s getting Wallace’s Line wrong, still lots of room for improvements as we tweak the model and the iNat training dataset grows
iNat keeps transforming into something better!
This looks really cool! Of course, my first instinct is to feed it information that will mess it up. What happens for very problematic, messy taxa such as Agaricus bisporus (most of the observations of which, in North America, are incorrectly identified)? @fungee
This feature should be very helpful for a lot of other fungi, I would expect.
Great work - this is such an amazing tool and the continued work to improve it is always appreciated and so helpful for future work!
Wow, this is amazing! I hope it eventually gets integrated into the observation and species maps more fully and as others have mentioned, gets more variable imputed. Perhaps average precipitation and temperature if you could find a good map of that, maybe someday even geology. Also maybe you can make a search page or link them to the taxon page instead of requiring one to paste in the taxa number (or maybe you did and i missed it? )
@loarie Thanks!
This is super cool, amazing work! The improvement in iNat's CV is great of course, but I also love the potential for contributions for ecology and conservation knowledge.
Using elevation is a clever way to estimate with more precision, like @pfau_tarleton and @gcwarbler and related to @brennafarrell's question this makes me curious about other variables for doing this. E.g. my immediate thought was of the Köppen climate classification, I'm not sure if that's based exclusively on precipitation and temperature or if vegetation data is including in its mapping? Presumably vegetation data would be more directly correlated with biodiversity than elevation. I also assume the global map is less precise for that than for elevation. But it sounds like the model is already incorporating other species' observations which is really cool.
Does the model include knowledge of oceanic elevation? I'm curious about its abilities to model pelagic species given their observations are going to be really biased towards coastal occurrences. Although I'm not sure how accurately we could judge its results given nobody knows a whole lot about where pelagic species spend their time.
Cool stuff. Two questions came to mind, one of which @jdmore already asked:
The other, re:
Really looking forward to that. Does the
original_geo_scoreproperty in the /v1/computervision/scoreobservation response contain the Geomodel score, or the 1-degree grid model score? If the latter, can/will the Geomodel score be added to the response as a separate property?@charlie.there is a link on the taxon page - see fig below “ we’ve linked from the taxon pages of all approximately included 80,000 species.” above
@upupa-epops the ocean is just set to one value in the elevation map we’re using, there’s no bathymetry data which would be best but I think the main constraint in oceans is so little inat data relative to on land so the model is mostly guessing
It doesn't seem to understand marine organisms very well. It predicts this strictly coastal fish lives mostly in the open ocean:
https://www.inaturalist.org/geo_model/49265/explain
Oh no :( Not like this: https://i.imgur.com/ETcQVJO.png
Not like this!
That's great!
Could you please review the wording wrt. percent and percent point (https://en.wikipedia.org/wiki/Percentage_point). Else the statement about a 4% increase in accuracy is incorrect, it's a 4pp increase.
Congrats for publishing this new models. Is it possible that it will act as a confirmation bias when used by non-expert users?
Geology would be an interesting extra layer. We have species found on granite, or sand, acid or lime.
So exciting to see big changes landing on iNat!
Top 1 suggestion from 83% to 87% - that's a big improvement given the room that was still there. Thanks for picking this up and making it a success.
oh, I'd be thrilled to be able to look at a subset of marginal / unusual observations. Talk about interesting! Whether an opportunity to correct a mistake or verify a cool new finding... I eagerly await further updates.
Awesome work! Thanks!
@dianastuder - wishful thinking.
Have you looked at the Cape data?: try https://www.inaturalist.org/geo_model/566478/explain (Ld salignum: the most catholic Fynbos plant species)
The entire Cape Flora is about 50 cells. At least 10 of these cells go from 200m to 1500+m altitude.
Just the Peninsula cell has granite, shale, sandstone, acid sands, alkaline sands: 5 major geological and more vegetational types - Forest, Sand Fynbos, Mountain Fynbos, Shale Fynbos, Granite Fynbos, Renosterveld. Strandveld.
The fine scale of biodiversity patterns in the Cape are invisible at this scale.
The really scary thing is that in these 50 cells are 20% (9000) of all plant species in Africa, and 70% occur ONLY in this area.
I asked earlier how many species are confined to a single cell?: I suspect that we might end up with over 1000 single-cell plant species in the Cape, although only a few dozen of these have over 100 observations at this stage.
Agree with dianastuder. I am a geologist by profession, and also feel that could be of some benefit.
Yes! Yes! Yes!
I am a botanist/ecologist by profession and I totally support the inclusion of geology - all our vegetation types are primarily determined by geology. While it will be useful in many parts of the world, in @dianastuder 's backyard it will be meaningless at this scale - we would need a grid at least 5-10 times finer before geological patterns will manifest in the Cape.
Excellent! Is there any plan in the future to make the unthresholded maps downloadable?
As tonyrebelo correctly says, it would have to be at least at a 1:1 000 000 scale for localised areas, which would probably be a mammoth undertaking.
I can provide that map of SA, and the soil map, if anybody is interested. rtb@ctecg.co.za
And for Zimbabwe, Botswana, and Namibia
Computer Vision should only work when the locality is already established. It would probaly prevent a lot of misidentified observations. Right now it is possible to first get the identification and to add the locality later when uploading an observation.
Currently, the 'CV included' label on taxa pages is only being displayed on a species level. Thus, the link to the geomodel for a specific taxon is not easily accessable (i.e. without knowing the link and taxon ID) for any higher taxon level. Would be nice to have this added.
I have just uploaded some observations with the new model. I am afraid that the geomodel has a bit too big "weight". When IDing liverworts, which are not a popular observation in my area (I live in a "liverwort - black spot"), my own previous findings spoil the suggestions. Before the new geomodel, Pellia species were perfectly recognized by the CV. Now the CV suggest Blassia pusilla instead of Pellia.
@bagous not putting in the location before asking CV, is a user choice (or error, depending on your own interpretation)
Unless it is Not Wild, when Visually Similar is more useful.
@raylantalbot I did not mean jpeg, tiff or any kind of image file format, but a geographic data file formats like GeoJSON or shapefile, which would have a reasonable size.
Very cool! Will this data be available via the API? For example getting a list of expected species at a location, or finding observations outside of where the geomodel expects? Could be useful in developing tools to help identify and fill data gaps.
Yes, it would be useful, but note the pixel size: the hexagon units are about 55km across.
Is this the resolution of computation, or just a safe summary level for sensitive species?
YESSSSSSSSSSSSSSSSS
@jujurenoult yes I perfectly accept that. I simply made the offer should anyone find a separate map useful in the meantime
"All models are wrong, but some are useful" - George Box. I'm excited that this appears to be a useful model for the intent it was designed for, and looking forward to future tweaks to make it even more useful! I'd love to see this incorporate other open data sources, if and when possible, such as those provided by other contributors to GBIF. Great job!
Congratulations, looks like you are heading in the right direction.
However, in terms of resolution, if from a continental point of view it is suitable for many situations, in others (including islands, locations with pronounced orography that enhances multiple habitats, etc.) the resolution is clearly insufficient. It seems it still can't effectively separate from a spatial point of view the different distribution between echium candicans and echium nervosum (mere example.)
I hope you continue to follow this path and that one day everything will be more precise/finer.
This now loeads to Rubus chamaemorus to not be suggested as Expected nearby even when there are many RG observations in the close vicinity:
This seems like a great improvement and as someone who studied biogeography I was hoping to look at the maps referenced here :"You can explore the Expected Nearby Maps and Unthresholded Maps we use to weight Computer vision suggestions on new Geomodel prediction pages we’ve linked from the taxon pages of all approximately included 80,000 species"
I have looked through all tabs on several taxa that have computer vision models but can see no link to their specific geomodel prediction pages (I'm especially interested in seeing the unthresholded maps). I was expecting to find it in the lower right corner of the about page where talks about computer vision model.
Can someone please tell me exactly where to look for these links on the taxon page for each species as I am just not seeing it?
Thanks, Tom
@taroman - on the species page, on the about tab, bottom right - where you suggested - click on the blue.
The "Expected Nearby" label is derived from the Geomodel.
Learn more about the Geomodel here.Could you be looking at species not yet in the Geomodel?
It is on the taxon page below the CV section.
Incredible work. I look forward to reading the papers on how these models were derived.
Is there a way to search for outliers to help with vetting? Thank you!
This is super cool and super needed work, thanks for the detailed explanation. I really enjoy your tech blog posts !
Hope this will apply to the work I am doing here on the recording and identification of species endemic to Haiti.
This is awesome! I can't wait until the "3. Future direction: Surfacing unusual observations" part is implemented! Are you planning to add a checkbox for these in Identify? That would be extremely helpful. As with gcwarbler's request, I'd be happy to evaluate unusual Euphorbia sect. Anisophyllum observations.
This is so cool.
The analysis appears to be extremely biased towards animals when only 9 out of than 4748 taxons are plants.
Making Inat more and more entertaining.
Thank you and congratulations!
@gcwarbler @loarie Global ecoregion GIS delineations are publicly available courtesy of The Nature Conservancy. I highly recommend training the geomodel with ecoregions, at least as a test. I believe the results will be exceptional.
https://tnc.maps.arcgis.com/home/item.html?id=7b7fb9d945544d41b3e7a91494c42930
Having now examined a few of the predictions of the GeoModel I'm a little less excited than I originally was.
For example the map for Trichocereus chiloensis shows regions east of the Andes with relative probabilities greater than threshold when in fact the species is endemic to Chile. I thought this might be a consequence of elevation being included in the model inputs but having read the paper I can't find a statement confirming that. Hopefully someone can enlighten me.
Awesome! I've been having a lot of fun playing around with the Geomodel.
I noticed that a link to the Geomodel isn't available for taxa above species, even if they are included in the CV (example: genus Spirogyra). I think it would be useful to provide a link in the "About" section for those as well!
I've looked at a few Euphorbia maps and feel that this still could use some development. It's not too bad, but seems to have difficulty dealing with range edges and disjunct populations. For instance, the model for Euphorbia capitellata over-predicts the range to the west and under-predicts to the east. Notably to the east, it doesn't predict where the occurrences actually are and instead predicts an area between the occurrences. From looking at the taxon range comparisons, it looks like recall could perform worse for plants than for animals.
@loarie You mention that adding additional covariates other than elevation didn't improve the models. What was the taxon sampling for this? In particular, were these compared for all groups separately or all groups collectively? Also, did it include uncommon and common taxa? This seems to do a little bit better for the more widespread Euphorbias.
Also, is there a way to add taxon ranges without relying on IUCN for maps? It seems like plants are getting the short end of the stick here because their distributions aren't available.
This is awesome! For its purpose (to help refine the visual recognition model) this seems like a resounding success already -- it having a tendency to include too many areas within the "expected" range is not a huge deal IMO since the goal is not to produce proper range maps. Awesome work.
@tonyrebelo, Thanks! I did see that but the way it was worded I thought it was just a link back to this page (guess I never tested it). I was expecting something that said "view species geomodel maps" or something like that!
Definitely seems like a step in the right direction, but sure could use some more geographic data parameters (such as water bodies) included. For instance in salamanders where ranges are often limited by major rivers the model will suggest expected in areas they have never occurred.
One of my favorites the Yonahlossee salamander does not exist west of the French Broad River, but both models extend it all the way through the Blue Ridge mountains west of the French Broad. I also wonder if my observations influenced the model as I am lead observer, but almost all my observations are from a private preserve in the SW corner of the range, close to this river (I obscure location, but assume the computer uses the exact locations). https://www.inaturalist.org/taxa/27224-Plethodon-yonahlossee
Another complication in salamanders is cryptic species, where the location is actually the deciding factor for species ID. Another of my favorites is the Blue Ridge Gray Cheeked salamander. The range map shown is quite accurate, but geomodel is way broader. While additional geographic parameters would likely improve this one, I'm sure there is only so much that can be done with cryptic species other than hard wiring the range map into the model (I'm surprised they don't have some special notation on species page for cryptic species). Of course I also wonder how much my data affects the model as I have about 80% of the observations & they are from one small private preserve. https://www.inaturalist.org/taxa/27172-Plethodon-amplus
Anyway as a supporter I am excited to see where this geomodel goes. Inclusion of elevation is a great start. Look forward to inclusion of more geographic parameters & better predcitions. But of course part of the beauty of nature is the uniqueness of species & kind of glad that no model could ever accurately predict where a species might be found!
Appreciate the explanations for this model. I naturalist is so useful & knowing how it works & its limitations makes it more so!
It's good that the CV (I'd still prefer to call it AI) ID suggestions are continuing to be gradually improved over time. Athough, the current CV still misidentifies many photos of cryptic taxa, like wasps and bees. Also, the Seek mobile app version doesn't use Nearby taxa at all in making suggestions, and for example misidentifies western honeybees in the US as asian honeybees every day. My only other feedback is about the following section from this post: "The change in name from Seen Nearby to Expected Nearby is intended to make it clear that the label comes from a model prediction rather than a grid of observations. Note that these predictions aren’t perfect."
Why not call it Predicted Nearby? Or Estimated Nearby or Potentially Nearby, or something like We Think is Nearby, which is used in other CV text, e.g., "We think this is X" and "We're not sure," which I'm paraphrasing. Many people will misinterpret Expected Nearby to mean the same thing as Seen Nearby. The most important issue is whether people reading that text will realize that the suggestions can be inaccurate, including out of range species in some cases. Seen Nearby and Expected Nearby wouldn't indicate that to many people, which is why I suggest Predicted, Estimated, or Potentially Nearby. It should also be considered that many people are observing and identifying without ever reading explanatory Help documents or how any of the models work, or who are only using mobile app versions with limited information, like the iOS app. I've met people who were only using the mobile version who even thought every CV suggestion was accurate.
"misidentifies western honeybees in the US as asian honeybees every day"
This is extremely tiresome and quite burdensome for identifiers.
Is it possible for the records flagged as outliers (at least the extreme outliers) by the models to be resubmitted as "in need of (further) identification" by identifiers? If reconfirmed as correct by identifiers (preferably by reliable ones by objective metrics) then the record can be used in future models, but if rejected or not confirmed maybe it shouldn't be?
When checking or rechecking records I would prefer if there were in addition to agree/disagree more nuanced additional options namely "not certain but likely correct" (may be acceptable if consistent with modeling results) or "not certain but likely wrong" (esp. suspect if inconsistent with overall modeling result).
Rechecking of outliers as revealed by modeling is very important in my opinion and should be rewarded if/when the data are used by non-parasitic analyzers for publications and other research projects.
Amazing work with incredible results
Replying to the current second to last comment. Re: outlier records, I also happened to bring up the issue of outliers that originate from non-inaturalist GBIF data-submitting sources earlier today https://github.com/gbif/portal-feedback/issues/4987. To address an issue that commonly occurs on inaturalist, where it's difficult to estimate how reliable a species outlier data point shown on GBIF is. On the one hand they could be valuable new locality records, and on the other they could be fake and waste identifier time.
Can we, do we, have a Best Practice for Outliers?
I leave a comment, and a broader ID - or use the CV for Seen Nearby if I am convinced. Their outlier and my good intentions will equal a broader CID either way.
For identifiers, the easiest would be to flag as 'outlier' on the taxon page. But that may not be useful to curators or the relevant iNat staff. An Outlier Project? Or a new Annotation?
Might the Geomodel Map become a map overlay in future? Similar to GBIF - so we can toggle on and off.
Any chance of having the Geomodel Map as an "Order" option in the Compare Tool and the Identotron? - in many of our regions the Place options are so large that the default order (number of records decending) does not help much at all.
@taroman I agree that this seems to be the most challenging scenario for this type of model: species that are constrained by geographical features (river, salt water, mountain range, whatever) from occurring in habitats that are otherwise suitable. This is a challenge for many ENM type models. Based on my understanding of this model, since it relies on training with the rest of the community around it - if there are species that are mobile and not subject to the same geographic barriers, then the model will overpredict and think that the focal species should also be present where the rest of the community does occur.
For salamanders restricted by drainage, the same tree, mammal, bird, plant species communities will often occur in nearby drainages, so the model will overpredict.
This situation might be a bit exacerbated here, because the species that are most commonly observed and most likely to be in the Geomodel are those more common species that have larger ranges and are capable of moving/dispersing more. The Geomodel view of communities in the "model space" is likely to be biased towards a broader community that is more homogenous - therefore I'd expect a broad pattern of overprediction in these cases.
This isn't necessarily a bad thing in some ways. For one, the most common, broad species are also most observed, so the model should work well for the vast majority of observation uploads of those most common species. For another, I'd rather have the model offer more suggestions for out of range stuff (which might be there) than underpredicting and being too cautious.
I do wonder if it might be possible to reduce some of the issues with this by adding information about the distance to existing observations of a taxon to the suggestions. With this approach, a suggestion for a given species could be downweighted the farther it is from any observation for that species in the training set. This approach seems like it could be any easy way to combine the information from the previous approach (which was purely spatially based) to this community-based predictive approach. This type of weighting usually isn't too hard to do (I don't think), and would be possible for all species included in the model, so it might be possible.
@cthawley, Thanks that does makes sense. Distance to other observations does seem like a good addition to the model, hope they can find a way to incorporate that & see if it helps! I wonder if they do or could weight taxonomic similarity in the model? Not perfect but there should be more similarity in mobility & dispersal abilities for more closely related taxa. For instance with salamanders around where I live in North Carolina (US), I think the majority of taxa have their ranges limited by the major river around here (The French Broad).
Thank you! It is a great step!
//Here is the traditional Chinese version: https://taiwan.inaturalist.org/blog/84743-geomodel
Just playing with the model, for species largely confined to a single cell (see https://www.inaturalist.org/projects/greater-cape-floristic-region/journal/84752-the-inaturalist-geomodel), a recurring pattern appears to be that the threshold is set just a tad too high, so that the cell with most records falls below the threshold, but a few neighbouring cells that are without any records are above the threshold. The model thus fails to predict the species within its distribution cell, but predicts it outside of its known range.
e.g. Snow Protea https://www.inaturalist.org/geo_model/592510/explain & Toothleaf Sugarbush https://www.inaturalist.org/geo_model/574546/explain
Failure to predict a species in 99% of its recorded (on iNat) range, seems to be a serious error.
A check on the threshold should catch this.
@tonyrebelo, interesting! This also holds for this salamander species where in the unthresholded model the highest probability areas are on the other side of the river which is a barrier to its distribution. & most records are from the same cell as this is a cryptic species with limited range. https://www.inaturalist.org/taxa/27172-Plethodon-amplus
I'm extremely excited about this being small enough for use on Seek and the iNaturalist app! It's much more convenient to snap a picture of a neat organism on my phone, but up until this point it ended up being more difficult if I didn't know what species I was photographing. How cool!
@loarie I just want to voice very strongly that removing captive observations from the model would be a big mistake. The fact that the previous "seen nearby" feature did not include captive observations led to huge numbers of misidentified observations in places they don't occur. For example, Quercus kelloggii, which the computer vision loved predicting in low elevation areas of California, despite the fact that it occurs only in mountains. This happens because the model is "forbidden" from looking at the correct species, Q. rubra, because nearby observations have all been marked captive. It requires a large amount of IDer effort to keep these incorrect IDs from polluting the data. There are many other plant species where this was an issue as well--basically anything where a similar species used in horticulture occurs somewhat near a similar looking wild species.
So the the change I'm actually most excited about from all this is that this problem shouldn't occur anymore!
Surely this argument should be extended to include hybrids, hybrid cultivars and especially invasive hybrids as well?
Exciting, though from comments n findings above seems there is lot room for improvement
Перевод на русский / Russian translation: https://www.inaturalist.org/posts/84807
Great research project!😍
This is wonderful! Happy to see improvements such as these.
this is awesome. nice to see Auchenorrhyncha representation in this post heh.
Another question of "Why would the geomodel predict this?" - I was just uploading an Artemisia observation from Ontario, Canada and the model oddly suggested A. pycnocephala, a Pacific coast species from Oregon/California not found within 3000km of the plotted location. The species doesn't show up as suggested on the geomodel map nor has any iNat observations nearby. Not sure why the geomodel would suggest it as expected. Overall the model seems like a great feature to improve accuracy, really fantastic to see elevation now being considered. Can't wait to see how it's further developed over time and expanded.
@cgbc that doesn't seem right - looking into it
I can't concur enough with @dianastuder that geology would be a fantastic addition in the future, particularly for geological types with a high rate of edaphic (soil-determined) endemism in plants - areas with a substrate of gypsum, limestone, dry sand, etc., and most of all, ultramafics/serpentinite. It would probably be easier and more useful to restrict incorporating surficial geology into the model to areas where the geology is likely to be a significant influence on the flora, rather than applying it worldwide with all rock types.
Macrostrat would be a good source to pull data from if incorporated. https://macrostrat.org/map/
Nice. I was dreaming of it (and proposed it) for many years in different citizen science projects developed in Europe (Faune France / Biolovision ; Orchisauvage,...). You did it! Congratulations.
Thanks for this exciting update, thrilled to see altitude being considered in the model. Also appreciated the fact that the range extension and mis-identification aspects are considered. I have a small question here, are the number of misdentifications also considered in the predictions somewhere and will it help improve the predictions, such as correlating misidentifications with geomodel? Or the species that is frequently misidentifed? It would help if the species with recurring misidentifications can be predicated more carefully.
The geomodel is a great addition! I'm glad to see that iNat was able to incorporate an elevation model as part of this. I would love for (assumed) elevation to be exposed as a search parameter for identification purposes. ;-)
I wonder whether there might be any benefit to using different threshold values for different high-level taxa groupings. It's fairly common for vagrant birds to show up far beyond their established range, whereas range extensions for well-studied plant species are less common and typically less extreme. Of course, there's a non-zero probability of almost any species in almost any location, but it seems that (pseudo-)probability threshold is the primary tool to achieve a good balance of suggestions that offers "just enough" improbability.
This has been at the top of my iNat wishlist for a while, excited to see this play out!
@cgbc we noticed a small bug in the Expected Nearby Map display that was rounding very small thresholds which explains why the Expected Nearby label for Coastal Sagewort you were seeing wasn't matching the Expected Nearby Map. That should be fixed now.
The Expected Nearby map for Coastal Sagewort of course now looks alot worse with a too liberal threshold. We experimented with a slight tweak to how we're generating thresholds. Its very easy to choose more conservative thresholds and get a better for for this species:


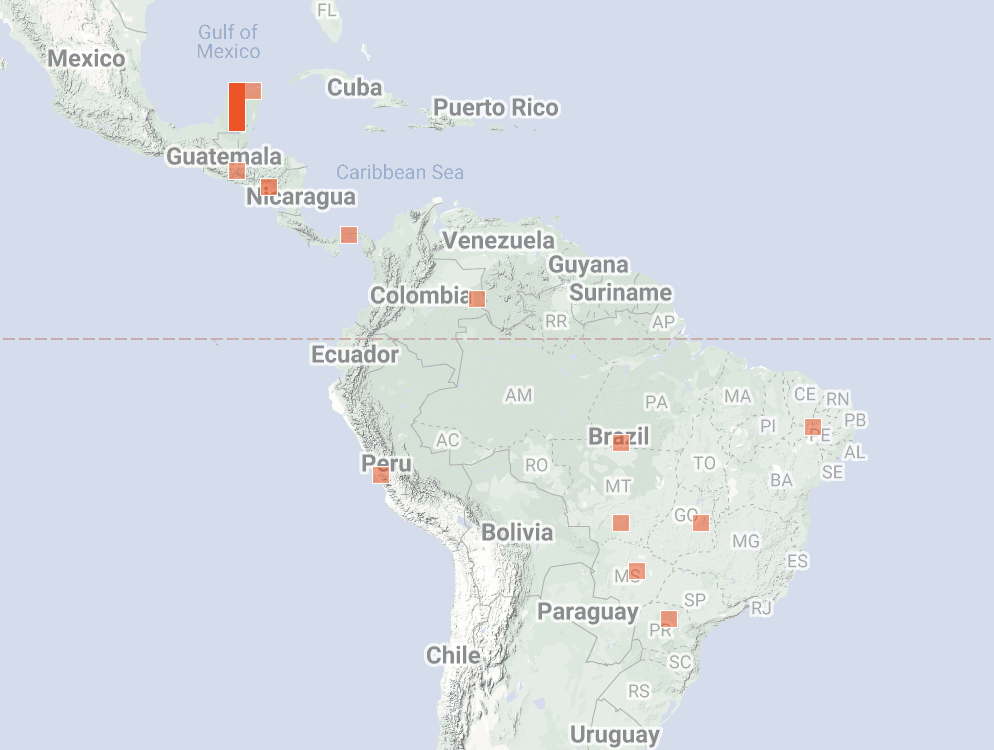
But these didn't evaluate any better on the taxon ranges overall - essentially did better on species like Coastal Sagewort where there were some (since fixed) misIDs out in Europe that really confused the model (what grows in only CA and Denmark??) and made the threshold too liberal
but it did worse on species like Broad-eared Free-tailed Bat by excluding legit points from the Amazon and further restricting the Expected Nearby prediction to the Yucatan driving the threshold too conservative
we'll continue to experiment with how to improve the model and threshold selection to hopefully find changes that improve the evaluation overall. In the meantime it looks like our very bad threshold for Coastal Sagewort is driven by since corrected misIDs from Europe, so at least we know that fixing misIDs is one way we know we can improve our threshold selection even if there is a lag between model versions before seeing the pay off.
Go guys Go!!!
What parameters are used as input for the Geomodel, also the datetime of the observation?
https://forum.inaturalist.org/t/introducing-the-inaturalist-geomodel/45407/15
Would it be useful to share examples of "Seen nearby" not working as expected anymore, presumably due to the Geomodel (it seems like it's totally ignoring some taxa if they're on the edge of their range now, even if they're locally common), or should I make a bug report for that (I have some screenshots if they're useful)?
E.g., it only suggested the antelope squrrel species from the nearby mountains for this observation, and not the common lowland species: https://www.inaturalist.org/observations/185010825
And for this observation https://www.inaturalist.org/observations/185010907 it didn't suggest Canada jays as nearby despite there being dozens of observations at this same parking lot.
?? cutoff threshhold a tad too high?
A bit like https://www.inaturalist.org/geo_model/574546/explain - where the threshold cuts off the only cell where the species actually occurs and has it only occurring outside of its natural range.
It looks like the issue I'm seeing is because of how the hexagonal cells in the Geomodel fall on the map. The spots where the two species I observed (Canada jays and white-tailed antelope squirrels) are falling a hundred meters or so outside a cell where the model does include those species, and so now it doesn't think they exist there at all. I feel like we'd at least want a gradient around the edge of the expected range, or a bigger buffer zone from existing observations on iNat.
Now I understand that elevation is included in the model used by iNaturalist I've been comparing the maps with those generated without that input. For the group I'm interested in (Cactaceae) it seems that elevation is, in some cases, actually detrimental to the output. Compare the two maps below and it's clear from the distribution of observations that Echinopsis leucantha should not be "Expected nearby" in Chile since it has never been observed there. The prediction obtained without elevation included (shown on the rhs) appears to better represent the species' distribution.
Based on the example @mrtnlowr found, it seems that using the current combination of elevation plus proximity of existing observations can be a poor predictor of likely locations in relation to mid-elevation species in mountain ranges. Many major ranges get the bulk of their precipitation from one direction (e.g. Andes, Himalaya, Sierra Nevada). Consequently, the middle elevations on one side of the range are relatively wet habitats, and on the other side (in the rain shadow) the same elevations are far drier.
The current model essentially assumes that locations west of the Andes at similar elevations to the known habitat of Echniopsis leucantha and 50–500 km from known observations are reasonable places to expect to find this species, despite the much wetter climate on the western slope and the existence of a 3800+ m crest in between. I'm not clear how much of this is determined by the parameters that the model is fed, and how much of it the model has learned on its own.
Obviously there's a balance to be struck here, and changes to narrow the scope of the geomodel might risk excluding likely locations for sparsely distributed species in terrain with less abrupt elevation and climate variation (e.g. the U.S. Midwest or Siberia). It would be interesting to see if running the optimization separately for, say, plants and birds (or for the "Southern Cone" vs. eastern North America) would produce better results. It seems that it would be theoretically possible to tailor geomodel predictions differently for a cactus, a warbler and a crab.
Very important for understanding how plate tectonics and continental drifts impacted evolution and distribution of species. Many common floral and faunal elements are found across continents now separated thousand of kms. Such models would help to understand biogeography, species evolution, island effect, geographical races, population and ecological genetics. This is very important development in biodiversity informatics
Very important for understanding how plate tectonics and continental drifts impacted evolution and distribution of species. Many common floral and faunal elements are found across continents now separated thousand of kms. Such models would help to understand biogeography, species evolution, island effect, geographical races, population and ecological genetics. This is very important development in biodiversity informatics
apologies if this has already been answered and I've missed it, but why do some species not have the expected nearby vs taxon range tab/map on their geomodel page? eg https://www.inaturalist.org/geo_model/566478/explain
Leucadendron salignum doesn't have a taxon range on the system at the moment so it doesn't have the taxon range overlay in the hamburger menu. As for the 'expected nearby vs taxon range tab', that appears if there is a taxon range in the system and if at least 90% of the training obs are within the taxon range (to minimize evaluating with taxon ranges with serious issues themselves).
right, makes sense, cheers
What sort of timeline is there for more broadly releasing a tool that helps identifiers locate outlier records? Since it was previously mentioned by @loarie that misidentified records have a negative impact on the geomodel, the sooner we can remove these points the better!
A suggestion for this tool, if we could combined the most unusual records from the geomodel with re-running the current CV model on these observations we could identify observations which are both geographically unusual, and the CV disagrees with. I've noticed that there are lots of observations of American species in Europe which were initally CV ids which are 2-3 years old, and now the CV has changed its mind with the most recent model and now suggests the correct ID. If we combine these tools we would have a very powerful approach for cleaning up some of the range maps which have been polluted with poor CV ids.
I like your suggestion @kevinfaccenda. It seems that some of the tools that @jeanphilippeb has produced, such as the series of "Unknown" projects, already have similar elements, such as utilizing CV ids as part of the process to highlight observations for human identifiers. Adding in some consideration of statistical outliers from the geomodel could really assist with this. In the meantime, of course we each can do a bunch of manual searches against the taxa where we have useful knowledge, but that seems a lot less efficient.
This is very interesting!
If I understand correctly, the model is built on lat, lon, and elevation, and does not include any other environmental variables. I think this means that it can't be used to predict how ranges will change under different climate change scenarios?
That suggests it might be a useful data filter/cleaner to use prior to conducting a more conventional SDM, which can be used for climate projections.
(I appreciate climate projections are not your primary objective here, just curious)
Link https://www.inaturalist.org/geo_model/975853/explain
The most biased model seen by me at the moment. A hexagon with the majority of records is out of the suggested six hexagons. Just for the notes.
Yesterday, I posted an issue in a different thread,
https://forum.inaturalist.org/t/geomodel-issue-often-observed-plant-no-more-suggested/46028
but it might better fit in here.
Lepidium graminifolium is a south european species which was often recorded in the region west of Frankfurt/Germany (about 50°North, 8°East; elevation from 80 meters at the Rhine river to ca. 500 m). The old CV suggestions included it, but nowadays it fails, e.g. in observation https://www.inaturalist.org/observations/187475552
So I tested a few more "odd" cases, i.e. species which can be encountered here, but actually belong to a far away place.
When the "expected nearby" filter is used,
Lindernia dubia still works
https://www.inaturalist.org/observations/130142634
Odontites luteus fails now
https://www.inaturalist.org/observations/179722407
Jurinea cyanoides still works in South Hesse
https://www.inaturalist.org/observations/181784400
and still fails 30 km away in Mainz
https://www.inaturalist.org/observations/185163908
The Rotenfels area is the only place in Germany where Lacerta bilineata can be found (and it is popular with iNatters, so there many observations); suggestions fail now:
https://www.inaturalist.org/observations/125579047
Between Bad Kreuznach and Bad Münster is a spot known for Natrix tesselata which is only found in that area - it fails now:
https://www.inaturalist.org/observations/83386803
The village of Schlangenbad is known for its local population of Zamenis longissimus, CV fails now:
https://www.inaturalist.org/observations/114272652
Dysphania pumilio still works:
https://www.inaturalist.org/observations/131592476
Eriocheir sinensis still works:
https://www.inaturalist.org/observations/130933697
Currently, I do not see a pattern when a species is no more suggested vs. when a species is still suggested. But I will take more care of looking into the "not expected nearby" suggestions.
Typo: "As describe in" should be "As described in"
thanks - fixed!
haww thats so cool..I'm in love with Inat .. i was thiking just a suggestion why dont we integrate AI in the platform actually..would make the data more accesible ..just a suggestion
Good evening, @loarie.
According to the first paragraphs of this page, "the Geomodel is trained only on iNaturalist observations and an elevation map." It is unclear whether the observations used to train the Geomodel for a specific species are just observations of that particular species or observations of other geographically related species as well.
For more than a year I've been searching for a specific endangered plant in central Portugal and the knowledge about related species has been absolutely essential for that search. That species in particular is quite discreet and not easy to spot, but there are half a dozen species that usually coexist with that one and are easier to see. Whenever I see 3, 4, 5 of those related species, I slow down and look more carefully because the endangered species that I am looking for might be there - and more often than not, it really is there. To some extent, the distribution of those related species can be used as a proxy for the distribution of the species that I have been searching. I've been doing that intuitively with quite good results... a Deep Learning Model would do it much better than my intuition.
About the coarseness of the public data available... I will send you a private message explaining two loopholes in iNaturalist's geoprivacy. (I shall not explain in a public comment for obvious reasons.)
Best regards.
@mferreira: The approach you use to derive inferences from associated plant communities makes sense, and I've seen it used by other researchers. However, I'm almost certain that the iNaturalist Geomodel is entirely species-specific. To use associated species as part of the Geomodel would require access to a comprehensive database of associations between organisms. Possibly, iNat's own data could be used to generate such associations, but I'd expect that would imply a lot of untested assumptions. Even if that could all be worked through, I wonder how much better the results would be compared to a much simpler species specific model.
If the model were going to be extended to add other factors, I'd think that various measures of rainfall, temperature and sunlight might be easier to incorporate.
It is using all species in a region. It's using machine learning to identify what other species each species is associated with
@kevinfaccenda: My understanding is that the Geomodel is treating each species separately: "For species X, where do existing iNat observations plus elevation data and proximity indicate that we would most likely find other occurrences".
I think what @mferreira is describing is an explicit component of association between species: "Species X is frequently found in proximity to species A–J. Let's predict the likelihood of finding species X based (partly?) on observations of species A–J." I don't believe iNat is doing that.
@rupertclayton Your understanding of the the iNat Geomodel is imcomplete. It does indeed, as @kevinfaccenda pointed out, use all taxa within a region simultaneously to 'learn' a distribution. I've looked at the code in great detail and can confirm that each observation, no matter what species, is represented in the model by only 5 parameters: 2 for longitude, 2 for latitude and one for elevation. It does learn by association.
Thanks @mrtnlowr. Perhaps @loarie can write a little more about how this works, or point us to an existing write-up.
@rupertclayton The paper describing the model is open access and describes in detail how the model is constructed with several tests of it's capabilities.
@rupertclayton taking those meteorological parameters into account (and also geology / soil composition) would lead us to a whole new level of predictability for the distribution of plants... Then instead of saying "this species might occur here" we would start saying "this species should occur here: if we don't find it, perhaps something is preventing it from occupying its habitat". That could be very informative for conservation purposes.
What parameters are used as input for the Geomodel, also the datetime of the observation?
https://forum.inaturalist.org/t/introducing-the-inaturalist-geomodel/45407/15
for training: latitude, longitude from iNaturalist observations and elevations derived from those coordinates
for inference: latitude, longitude, elevation at a point
currently date is not being used
Are the data accessible to download? It would be great to be able to download GIS data from the Geomodel for our work.
not currently - but we would like to make the data available
Add a Comment