New validations FYI curators
Several parts of iNaturalist, such as the ‘community identification’ algorithm and ongoing efforts to make the computer vision algorithm work across the iNaturalist taxonomy (ie not just on species) require that active taxa grafted to the taxonomy tree have ancestries with only:
- coarsening ranks (e.g. if the taxon is of rank family, the parent can’t be rank genus)
- active taxa (e.g. if a taxon is active, its parent can’t be inactive)
To help discourage iNaturalist curators from accidentally introducing these kinds of inconsistencies, we’ve added a few new validations.
Using the Family Salamandridae as an example, curators can no longer do things like:
- change it’s rank to Subphlyum as this would violate the ancestries with coarsening ranks condition
- inactivate it as this would violate the ancestries with only active taxa condition
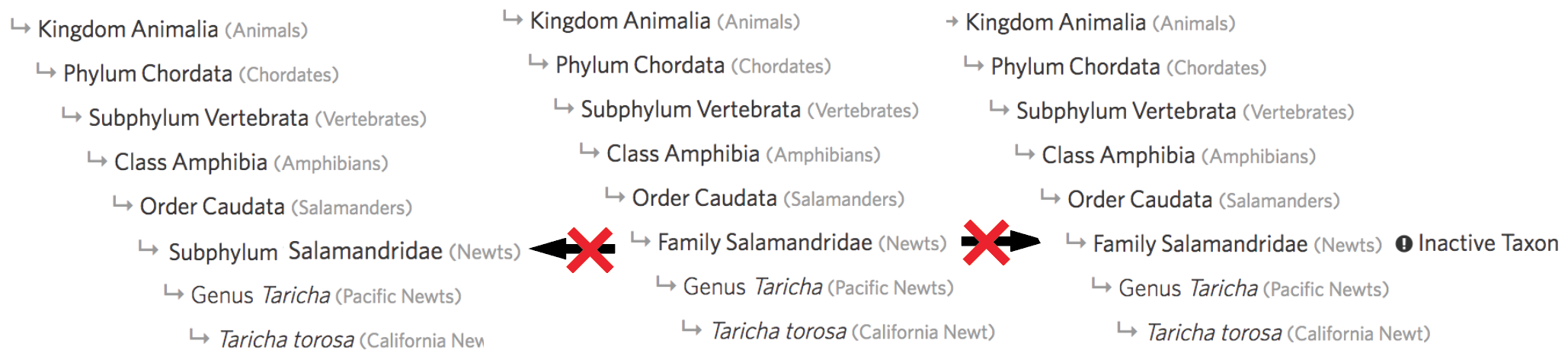
It was previously possible for curators make each of these edits to the taxonomy directly. However, the most common culprit for introducing rank inconsistencies was the automated process for importing names from external names providers like Catalog of Life which has also been reigned in by these validations. The most common culprit for introducing inactive taxa inconsistencies was through taxon changes. Imagine a draft taxon change structured as follows:
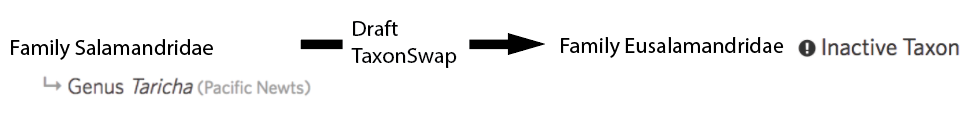
Committing a taxon change first inactivates the input taxa (e.g. Family Salamandridae) and then activates the output taxa (e.g. Family Eusalamandridae). In the past, committing taxon changes left children behind leaving situations like this:
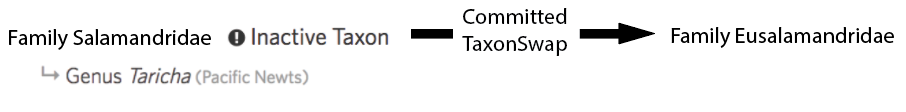
Which, unless curators take the time to manually handle the children after committing, resulted in active taxa lingering behind with inactive parents. This led us to make a change such that taxon changes with single outputs (ie Taxon Swaps and Taxon Merges but not Taxon Splits) automatically moved children like this:

Or, when the names of children depend on the parent (e.g. a species binomial), automatically created taxon swaps for the children like this:
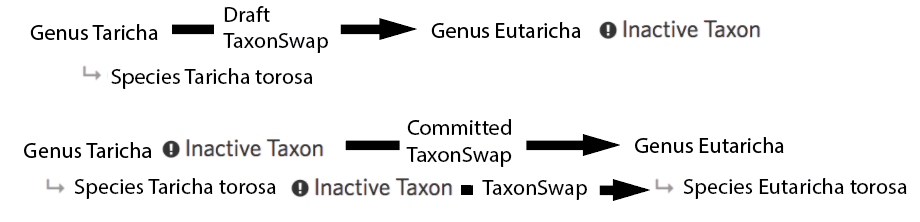
This approach to always automatically handling children has led to some screwy taxon names because it’s ignorant of things like gender agreement (e.g. Kyllinga alata moved to Cyperus should be Cyperus alatus, not Cyperus alata). And because it doesn’t work on taxon changes with multiple outputs (ie Taxon Splits) curators are still making taxon changes that ‘leave behind’ children.
As part of these new changes described here, we’ve made it optional to automatically move children on commit when making a Taxon Swap or Taxon Merge

But we’ve also made it so that iNaturalist will prevent you from committing a taxon change if the input taxon has active children and you haven’t checked ‘move children to output’. This is intended to encourage curators to handle all the active children before committing a taxon change. As before, you can handle this automatically by checking ‘move children to output’ for Swaps and Merges but please use it with care to avoid creating weird binomials like the gender issues described above. For Splits, you have to now handle the children first manually.
One annoying Catch-22 here is that these same new validations would prevent you from first moving an active child to the inactive output of the draft taxon change before committing, because active taxa can no longer be moved to inactive parents. Doh...
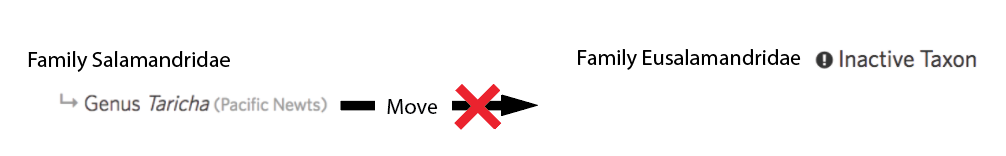
So we’ve added the caveat where you can move active taxa to inactive parents if the parents are the outputs of draft taxon changes:
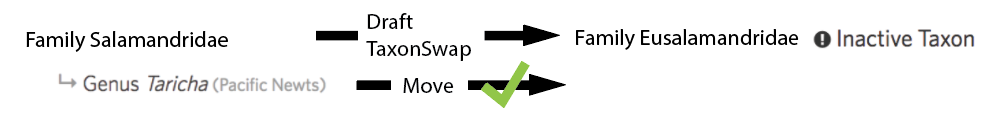
So remember to first make the draft taxon change, then handle the children manually (or by checking 'move children to output' when appropriate), and then finally commit the draft taxon change.
We recognize that these new validations might cause a bit more work for everyone upfront when curating taxa to make sure that the integrity of the ancestries isn’t corrupted. But we hope that this will create less cleanup for everyone in the long run. On that last note, a huge thanks to everyone who’s helped cleanup the hundreds of existing rank and inactive taxa issues already in the taxonomy over the last few weeks - especially @kokhuitan and @bouteloua who have done so much of this. All the rank issues are sorted and I’ve pasted links to the remaining 68 inactive taxa issues below in case you want to help. We’ll monitor to see whether these kind of inconsistencies are still getting introduced through pathways we haven’t considered yet.
Thanks again for all your help curating iNaturalist!
Mentioning our top 20 curators:
@treichard, @choess, @bouteloua, @borisb, @stephen_thorpe, @maxkirsch, @hkmoths, @kokhuitan, @duarte, @berkshirenaturalist, @jakob, @jonathan142, @cmcheatle, @tiggrx, @coreyjlange, @bobby23, @sea-kangaroo, @tonyrebelo, @eol_education, @kai_schablewski
FYI Remaining 68 inactive taxa with active descendants:
https://www.inaturalist.org/taxa/126338
https://www.inaturalist.org/taxa/160773
https://www.inaturalist.org/taxa/82890
https://www.inaturalist.org/taxa/366718
https://www.inaturalist.org/taxa/152909
https://www.inaturalist.org/taxa/142375
https://www.inaturalist.org/taxa/178397
https://www.inaturalist.org/taxa/246174
https://www.inaturalist.org/taxa/84009
https://www.inaturalist.org/taxa/173395
https://www.inaturalist.org/taxa/72208
https://www.inaturalist.org/taxa/142368
https://www.inaturalist.org/taxa/181363
https://www.inaturalist.org/taxa/148483
https://www.inaturalist.org/taxa/49148
https://www.inaturalist.org/taxa/164599
https://www.inaturalist.org/taxa/163781
https://www.inaturalist.org/taxa/221264
https://www.inaturalist.org/taxa/126870
https://www.inaturalist.org/taxa/60172
https://www.inaturalist.org/taxa/211511
https://www.inaturalist.org/taxa/424784
https://www.inaturalist.org/taxa/175776
https://www.inaturalist.org/taxa/335007
https://www.inaturalist.org/taxa/69580
https://www.inaturalist.org/taxa/208727
https://www.inaturalist.org/taxa/152861
https://www.inaturalist.org/taxa/61126
https://www.inaturalist.org/taxa/366574
https://www.inaturalist.org/taxa/211389
https://www.inaturalist.org/taxa/123845
https://www.inaturalist.org/taxa/176360
https://www.inaturalist.org/taxa/379251
https://www.inaturalist.org/taxa/82412
https://www.inaturalist.org/taxa/164817
https://www.inaturalist.org/taxa/71657
https://www.inaturalist.org/taxa/336176
https://www.inaturalist.org/taxa/244116
https://www.inaturalist.org/taxa/346633
https://www.inaturalist.org/taxa/77141
https://www.inaturalist.org/taxa/359516
https://www.inaturalist.org/taxa/59095
https://www.inaturalist.org/taxa/62354
https://www.inaturalist.org/taxa/152932
https://www.inaturalist.org/taxa/163573
https://www.inaturalist.org/taxa/372761
https://www.inaturalist.org/taxa/148975
https://www.inaturalist.org/taxa/343503
https://www.inaturalist.org/taxa/349315
https://www.inaturalist.org/taxa/321807
https://www.inaturalist.org/taxa/203833
https://www.inaturalist.org/taxa/157125
https://www.inaturalist.org/taxa/342574
https://www.inaturalist.org/taxa/347180
https://www.inaturalist.org/taxa/338640
https://www.inaturalist.org/taxa/320761
https://www.inaturalist.org/taxa/122111
https://www.inaturalist.org/taxa/246143
https://www.inaturalist.org/taxa/127138
https://www.inaturalist.org/taxa/324196
https://www.inaturalist.org/taxa/363465
https://www.inaturalist.org/taxa/359159
https://www.inaturalist.org/taxa/511731
https://www.inaturalist.org/taxa/355276
https://www.inaturalist.org/taxa/174140
https://www.inaturalist.org/taxa/118355
https://www.inaturalist.org/taxa/203106
https://www.inaturalist.org/taxa/117765





Comments
Usually, I don't like to interfere out of my taxonomic scope - I may reproduce errors, not being able to judge upon the reliability of web sources.
However, I picked one from the list above, and it seemed to be a clear case: https://www.inaturalist.org/taxon_changes/39770
I guess it should read "1. coarsening ranks (e.g. if the taxon is of rank FAMILY, the parent can’t be rank GENUS)"
First ten entries of above list clear!
thanks jakob - fixed
Another validation required? number of words in a taxon name.
see https://www.inaturalist.org/taxa/593894-Searsia-lucida - note the names. These were posted as species instead of forms.
Should the number of words in a name not be checked? Usually only one word in a name, but two for a species and three for an infraspecies. But names posted with four or more names should be flagged for checking.
(some names do require more words - genera and species of micro-organisms for instance)
Add a Comment